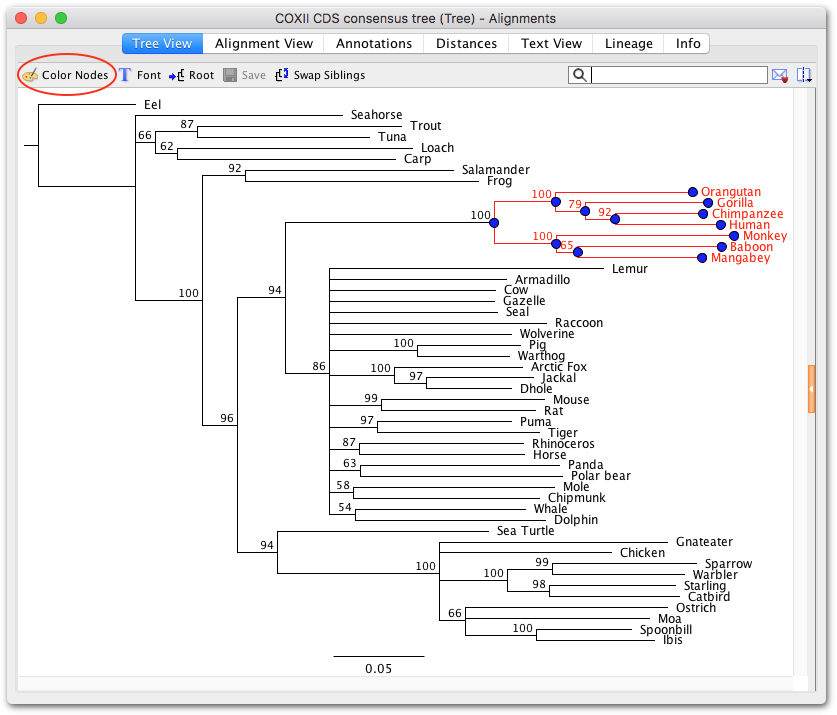
( A) Dot plot comparing homology of each poxin structure to representatives in the Protein Data Bank. Error bars are ± standard error of the mean for n = 3 independent experimental replicates. Note, the histidine inactivating mutation likely also impairs 2’3’-cGAMP resulting in an underestimated K D value. ( H) Quantification of VACV H17A and AcNPV H46A gel shifts in E and F, and determination of K D for 2′3′-cGAMP. Each image is representative of three independent experimental replicates. ni poxin using a range of 2′3′-cGAMP concentrations from 0.025 to 20 µM. No stable 2′3′-cGAMP binding was observed for mutant T. ( E–G) Gel shift analysis of stable poxin–2′3′-cGAMP complex formation using active-site mutant proteins. ni exhibits much higher K M and K cat values, and a lower catalytic efficiency. In contrast, the lepidopteran poxin from T. Viral poxin homologs from poxviruses and baculoviruses share similar enzyme kinetics and catalytic efficiencies. ( D) Table showing values computed from plots in ( A–C) giving K M, K cat, and catalytic efficiency for each tested enzyme with 95% confidence interval in parentheses below. Each plot is representative of at least two independent experiments. ( A–C) Plots of enzyme kinetics data for poxin enzymes from Groups 1, 2, and 3. Error bars are ± standard error of the mean for n = 3 technical replicates. ( G) Shared poxin active-site features (see Figure 2-figure supplement 1C) include hydrophobic pockets for each base of 2′3′-cGAMP and charged interactions that read out the 2′–5′ phosphodiester linkage. ( F) Despite differences in active-site residues ( Figure 2-figure supplement 1B,C), 2′3′-cGAMP is contorted within each structure into a similar strained conformation, confirming that poxins operate by the same core metal-independent mechanism established for VACV poxin. Consistent with alternative active-site residues, lepidopteran poxin enzymes show remarkably different kinetic properties ( Figure 2-figure supplement 2). ni and betabaculovirus PrGV poxin proteins contain altered active-site triads composed of two histidines, and an arginine or lysine residue. ( B, C) Conservation of the active-site triad between VACV and AcNPV poxin. ( A) X-ray crystal structures of poxin enzymes from an alphabaculovirus (AcNPV), the moth Trichoplusia ni, and a betabaculovirus (PrGV) allow comparison with VACV poxin (PDB: 6EA9) and demonstrate family-wide structural conservation ( Figure 2-figure supplement 1A). Data are representative of two independent experiments. See Materials and methods for protein accession numbers. coli (red text, empty lanes on TLC plate).  
Four proteins could not be expressed in E. Recombinant proteins ( Figure 1-figure supplement 1B) were incubated with radioactively labeled 2′3′-cGAMP for 1 hr at 37☌, and degradation products were resolved using thin-layer chromatography (TLC). Poxin 2′3′-cGAMP nuclease activity is conserved within each group. Poxins can be divided into groups, including enzymes from mammalian and insect poxviruses (Group 1), parasitoid wasps and alphabaculoviruses (Group 2), moths and butterflies (Lepidoptera) (Group 3), and cypoviruses, betabaculoviruses, and betaentomopoxviruses (Group 4). ( B) Bioinformatic identification ( Figure 1-figure supplement 1A) and biochemical verification of diverse poxin enzymes. Poxin enzymes inhibit cGAS-STING signaling by degrading 2′3′-cGAMP and blocking activation of STING. 2′3′-cGAMP activates the receptor Stimulator of Interferon Genes (STING) and initiates downstream antiviral signaling. ( A) Schematic of cGAS-STING signaling. The sensor cyclic GMP–AMP synthase (cGAS) detects cytosolic DNA and synthesizes the second messenger 2′3′-cGAMP from ATP and GTP.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
